GGPLOT LOG SCALE TRANSFORMATION
This article describes how to create a ggplot with a log scale. This can be done easily using the ggplot2 functions scale_x_continuous() and scale_y_continuous(), which make it possible to set log2 or log10 axis scale. An other possibility is the function scale_x_log10() and scale_y_log10(), which transform, respectively, the x and y axis scales into a log scale: base 10.
Note that, the scale functions transform the data. If you fit anything to the data it would probably change the fitted values.
An alternative is to use the function coord_trans() for transformed Cartesian coordinate system. coord_trans() is different to scale transformations in that it occurs after statistical transformation and will affect only the visual appearance of geoms.
In this R graphics tutorial, you will learn how to:
- Log transform x and y axes into log2 or log10 scale
- Show exponent after the logarithmic changes by formatting axis ticks mark labels.
- Display log scale ticks. R function: annotation_logticks()
Contents:
- Key ggplot2 R functions
- Set axis into log2 scale
- Set axis into log10 scale
- Display log scale ticks mark
- Conclusion
Key ggplot2 R functions
Start by creating a scatter plot using the cars data set:
library(ggplot2)
p <- ggplot(cars, aes(x = speed, y = dist)) +
geom_point()
p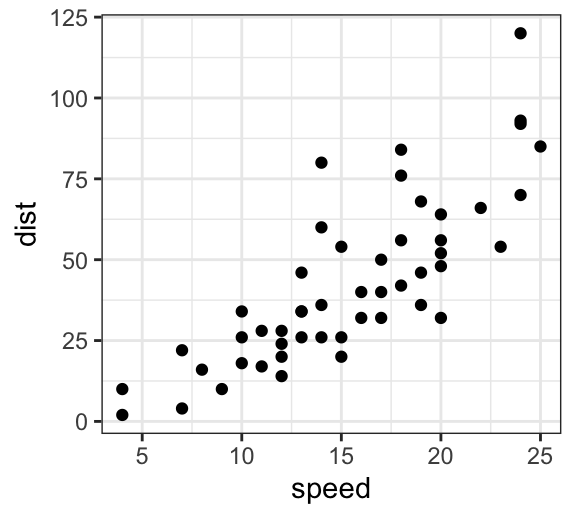
R functions to set a logarithmic axis:
- p + scale_x_log10(), p + scale_y_log10() : Plot x and y in log 10 scale, respectively.
- p + coord_trans(x = “log2”, y = “log2”): Transformed cartesian coordinate system. Possible values for x and y are “log2”, “log10”, “sqrt”, …
- p + scale_x_continuous(trans = “log2”), p + scale_y_continuous(trans = “log2”). Allowed value for the argument trans, include also ‘log10’.
- p + scale_y_log10() + annotation_logticks(): Display log scale ticks.
Set axis into log2 scale
- Log2 transformation of x and y axes
- Format ticks label to show exponents
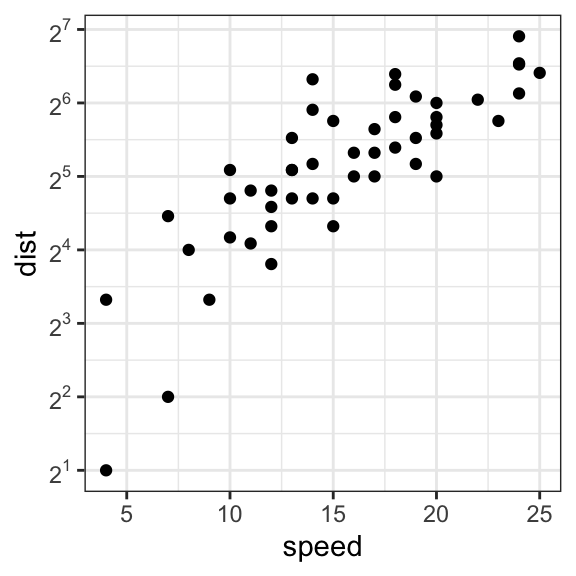
# Possible values for trans : 'log2', 'log10','sqrt'
p + scale_x_continuous(trans = 'log2') +
scale_y_continuous(trans = 'log2')
# Format y axis tick mark labels to show exponents
require(scales)
p + scale_y_continuous(trans = log2_trans(),
breaks = trans_breaks("log2", function(x) 2^x),
labels = trans_format("log2", math_format(2^.x)))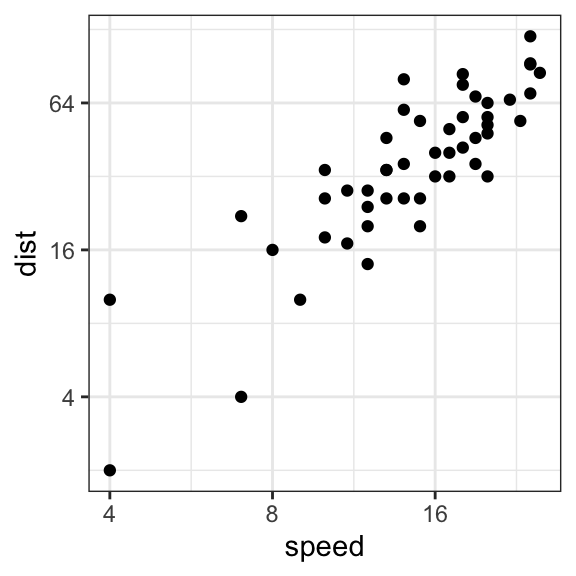
Set axis into log10 scale
The following R code changes the y axis scale into log10 scale using the function scale_y_log10():
p + scale_y_log10()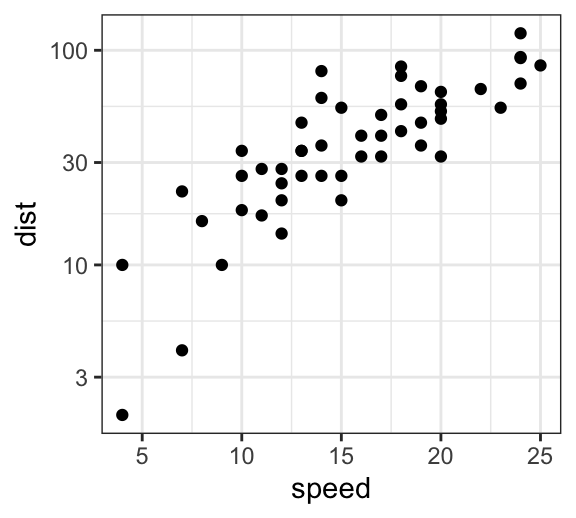
Alternatively, you can use the function scale_y_continuous(), which allows to transform breaks and the format of labels.
p + scale_y_continuous(trans = "log10")Display log scale ticks mark
Note that, log scale ticks make sense only for log scale base 10.
Key function: annotation_logticks() Data: Animals data sets, from the package MASS
- Create a log-log plot without log scale ticks:
# Load packages
require(MASS) # to access Animals data sets
require(scales) # to access break formatting functions
data(Animals) # load data
# x and y axis are transformed and formatted
p2 <- ggplot(Animals, aes(x = body, y = brain)) + geom_point() +
scale_x_log10(breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(10^.x))) +
scale_y_log10(breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(10^.x))) +
theme_bw()
p2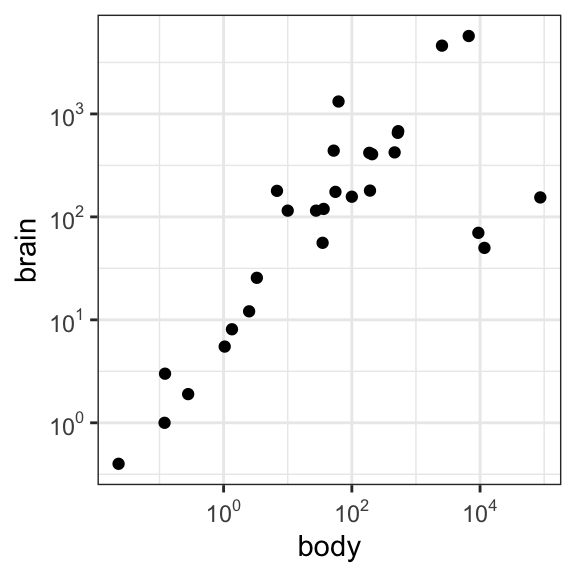
- Show log scale ticks:
p2 + annotation_logticks() 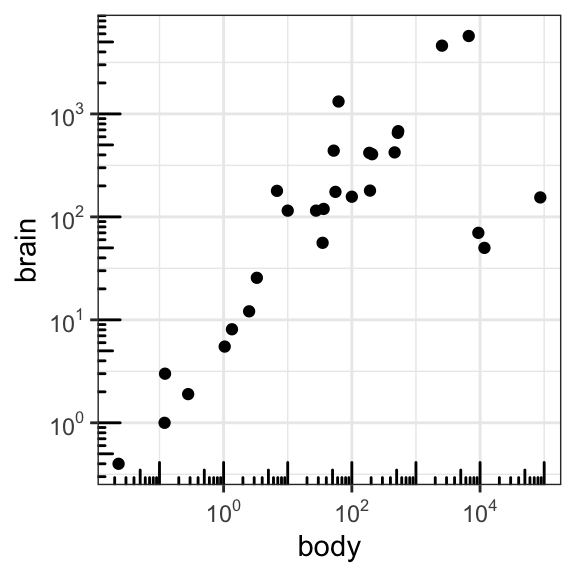
Note that, default log ticks are on bottom and left. To specify the sides of the log ticks :
# Log ticks on left and right
p2 + annotation_logticks(sides="lr")
# All sides
p2+annotation_logticks(sides="trbl")Allowed values for the argument sides are the combination of “t” (top), “r” (right), “b” (bottom), “l” (left).
Conclusion
We introduce how to create a ggplot with log scale. Briefly, the steps are as follow:
- Create an example of ggplot:
library(ggplot2)
p <- ggplot(cars, aes(x = speed, y = dist)) +
geom_point()- Log transformation of the axis scale:
# log base 2 scale
p + scale_x_continuous(trans = 'log2') +
scale_y_continuous(trans = 'log2')
# Log base 10 scale + log ticks (on left and bottom side)
p + scale_x_continuous(trans = 'log10') +
scale_y_continuous(trans = 'log10')+
annotation_logticks(sides="lb")
Python Example for Beginners
Two Machine Learning Fields
There are two sides to machine learning:
- Practical Machine Learning:This is about querying databases, cleaning data, writing scripts to transform data and gluing algorithm and libraries together and writing custom code to squeeze reliable answers from data to satisfy difficult and ill defined questions. It’s the mess of reality.
- Theoretical Machine Learning: This is about math and abstraction and idealized scenarios and limits and beauty and informing what is possible. It is a whole lot neater and cleaner and removed from the mess of reality.
Data Science Resources: Data Science Recipes and Applied Machine Learning Recipes
Introduction to Applied Machine Learning & Data Science for Beginners, Business Analysts, Students, Researchers and Freelancers with Python & R Codes @ Western Australian Center for Applied Machine Learning & Data Science (WACAMLDS) !!!
Latest end-to-end Learn by Coding Recipes in Project-Based Learning:
Applied Statistics with R for Beginners and Business Professionals
Data Science and Machine Learning Projects in Python: Tabular Data Analytics
Data Science and Machine Learning Projects in R: Tabular Data Analytics
Python Machine Learning & Data Science Recipes: Learn by Coding
R Machine Learning & Data Science Recipes: Learn by Coding
Comparing Different Machine Learning Algorithms in Python for Classification (FREE)
Disclaimer: The information and code presented within this recipe/tutorial is only for educational and coaching purposes for beginners and developers. Anyone can practice and apply the recipe/tutorial presented here, but the reader is taking full responsibility for his/her actions. The author (content curator) of this recipe (code / program) has made every effort to ensure the accuracy of the information was correct at time of publication. The author (content curator) does not assume and hereby disclaims any liability to any party for any loss, damage, or disruption caused by errors or omissions, whether such errors or omissions result from accident, negligence, or any other cause. The information presented here could also be found in public knowledge domains.
